Our groups
In this new era of large scale data analysis as well as rapid developments in the sequencing and analysis of the human genome, there is a need to take a multidisciplinary approach. If you are interested in genomics, big data analysis and digital health including personalized medicine, then this program is perfect for you. You will be interacting with a wide range of people and disciplines to advance personalized health and basic understanding of the variability of human biology.

Emergency and Disaster Pharmacy
Emergency and disaster pharmacy is a research field that focuses on ensuring safe, rapid and equitable access to medicines and medical devices during crises, such as natural disasters, humanitarian emergencies or conflicts. Notably, it addresses supply chain resilience, emergency drug protocols and patient safety in extreme conditions. This field also aims to assess the use of e-health tools, real-time data systems, telepharmacy and AI-driven logistics to support decision-making, optimise resource allocation and maintain the continuity of pharmaceutical care when traditional healthcare infrastructures are disrupted.
Apply now

Quality of care: patient-centered, efficient, equitable care
Research on quality of care examines how healthcare systems can provide patient-centered, efficient, and equitable care. Delphine Courvoisier's labs uses advanced methodologies (AI, LLM, longitudinal models) to explore how patient-level, healthcare provider level, relationship-level (e.g. shared decision-making), and healthcare level factors influence both hard outcomes and patient experiences and outcomes (PREM and PROM). Specifically, current research focus on inequalities in access to care and perceived healthcare quality, establishing quality register in rheumatic and musculoskeletal diseases, and the development of indicators to measure equity in hospitals.
Apply now

Genomics and genetics in evolutionary perspective
Genomics and genetics is transforming with advances of sequencing technologies. It also transforms the healthcare, from assessing human genetics, to tracking microbial outbreak or antibiotic resistance, and associating microbiomes with diseases. These big data challenge our computational techniques as well as our evolutionary models to interpret the genetic diversity.
Apply now
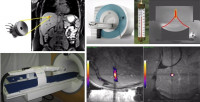
Biomedical Engineering and Interventional Imaging
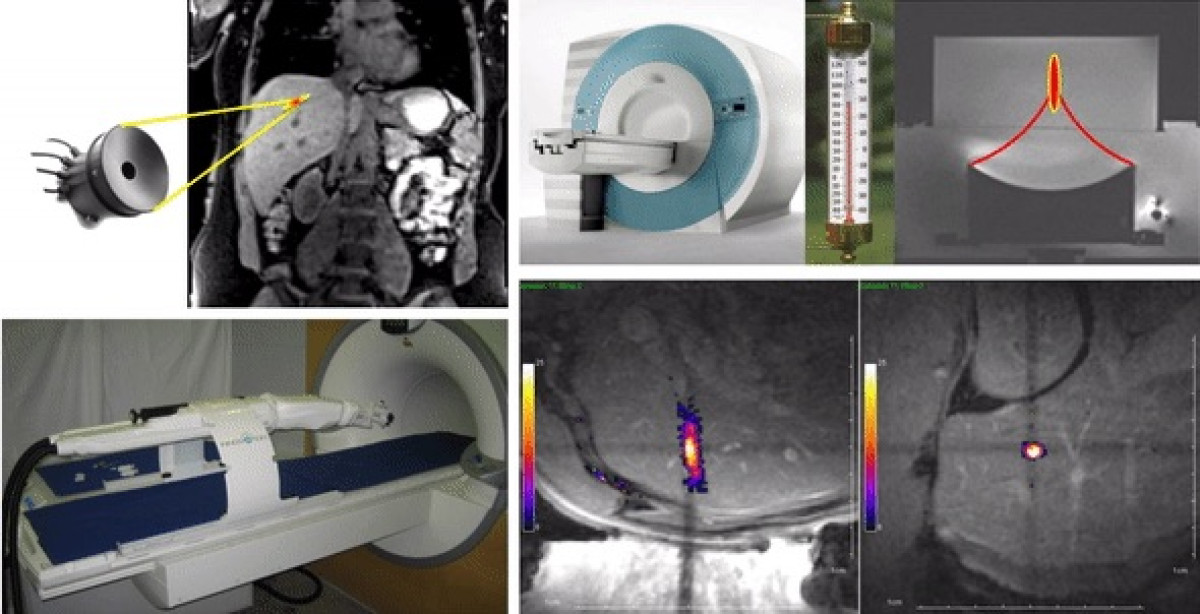
The work group comprises research scientists and radiologists. The team has a wide range of competences in biomedical engineering, interventional radiology, intra-operative image guidance, computer sciences. Our topics cover translational and clinical research, in particular in the field of oncology, from in-house prototyping of innovative devices to clinical trials using image-guided novel therapies. High intensity focused ultrasound (HIFU) is currently the only extra-corporeal technology capable of depositing sharply delineated energy patterns deep inside the body. Magnetic resonance imaging provides excellent capabilities for spatial guidance of HIFU therapy and is currently the only available technique for non-invasive temperature measurement within biological tissue.
Apply now

Data Science for Digital Health
The Data Science for Digital Health (DS4DH) group aims to design and implement novel algorithms and computational methods for the management and analysis of complex and large-scale datasets in the health and life sciences domain in order to foster innovative digital health solutions. The group is particularly interested in developing research around machine learning and natural language processing models that can blend and exploit (semantically) rich and (often) non-Euclidean datasets to create actionable insights. DS4DH is located at Campus Biotech and works closely with other digital health players, such as SIB, HUG and the SIMED, BiTeM and HI5lab groups. It is also well connected with private actors working at the forefront of innovation in the health and life sciences sectors.
Apply now

Epigenetic Control of Developmental Processes
Our aim is to understand how epigenetic information, in particular the 3D organization of human and mouse genomes instruct the development of organs and structures during embryogenesis. To map and functionally disrupt these chromatin states, we employ a variety of state-of-the-art technologies in pluripotent stem cells and in vivo during embryogenesis. The understanding of these basic concepts is critical to unravel the molecular mechanisms that underlie pathological gene misregulation in congenital malformations and cancer as well the mechanism that enable evolutionary novelties and diversity of life.
Apply now

Non-Compliance in Aesthetic Medicine: Public Health
At the core of public health and therapeutic governance, the research conducted by the Cantonal Pharmacist’s Office focuses on the safety, compliance, and proper use of medicines and medical devices to safeguard population health. Its research investigates determinants of non-compliance in aesthetic medicine, particularly regarding fillers and botulinum toxin, within nationwide campaigns. It also examines social media’s influence on product use and identifies social, behavioral, and regulatory factors contributing to unsafe practices. By collaborating with regulatory authorities, the office promotes evidence-based interventions to strengthen public health protection.
Apply now

Personalized Reconstructive Dentistry & Oro-Facial Rehabilitation Research Group
Our research group focuses on advancing personalized reconstructive dentistry and oro-facial rehabilitation through cutting-edge digital technologies. We integrate advanced 3D imaging, AI-supported treatment planning, and big data analytics to develop precision solutions for complex dental and facial reconstruction to enhance clinical outcomes. Our interdisciplinary approach reinforces collaboration across dentistry, medicine, and computational sciences, creating a dynamic environment for pioneering research. Key research areas include: 1. Development of AI-powered diagnostic and treatment planning tools; 2. Digital workflow optimization for personalized dental prosthetics; 3. Machine learning applications in patient outcome prediction; 4. Cross-disciplinary integration of dental, medical, and computational expertise. We offer PhD candidates the opportunity to work at the intersection of digital dentistry, computer science, and medicine, collaborating with specialists across multiple domains to transform patient care through technology-driven innovations.
Apply now

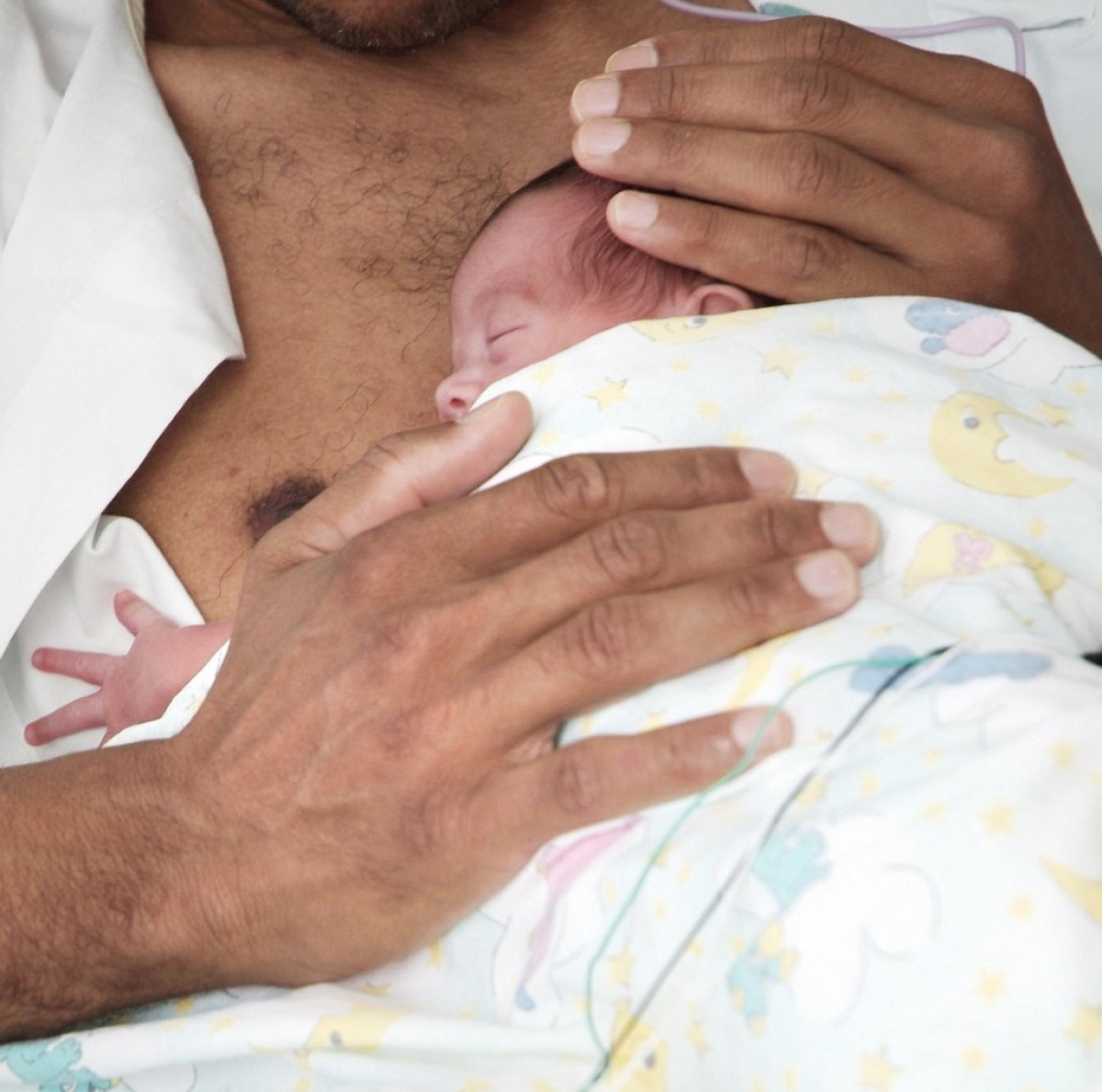
Digital Health Support for Neonatal Care
Over 15 million infants are born prematurely each year worldwide. In Switzerland, many require extended NICU stays, creating major emotional and practical challenges for families. Despite global recommendations, no accessible digital support tool exists for the transition from NICU to home. We will develop an mHealth app co-designed with parents, clinicians, and digital health experts. Features include evidence-based content, development tracking, a shared care calendar, secure messaging, and a digital memory album. Led by HUG and an interdisciplinary team, the project includes an acceptability study and RCT to assess usability and impact on parental stress, infant health, and healthcare use.
Apply now

Medical Information Sciences
Data, algorithms and knowledge for human’s health. Digitalization is a major change in our society and applies to the whole life science ecosystem. The “Medical Information Science” group is working on the phenotypic side. Working with very heterogeneous sources of multimodal data - personal health records, such as sensors, captors and activities, patients-related data, such as computerized patient records, behavior and lifestyle - environment, exposition factors - the living ecosystems - regulatory frameworks - knowledge sources in order to build actionable data pipelines that can be used to connect with *omics. The activities of the group focus on semantics, data interpretability tools such as natural language processing. People work with symbolic, rule-based and probabilistic instrument
Apply now
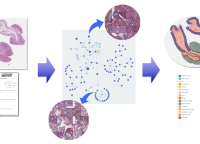
Multimodal & explainable machine learning
Our group works on approaches that combine machine learning on medical image data with text analysis approaches to build clinical decision support systems in radiology, histopathology and ophthalmology. A particularly focus is put on the interpretability of the machine learning models.
Apply now
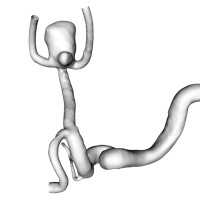
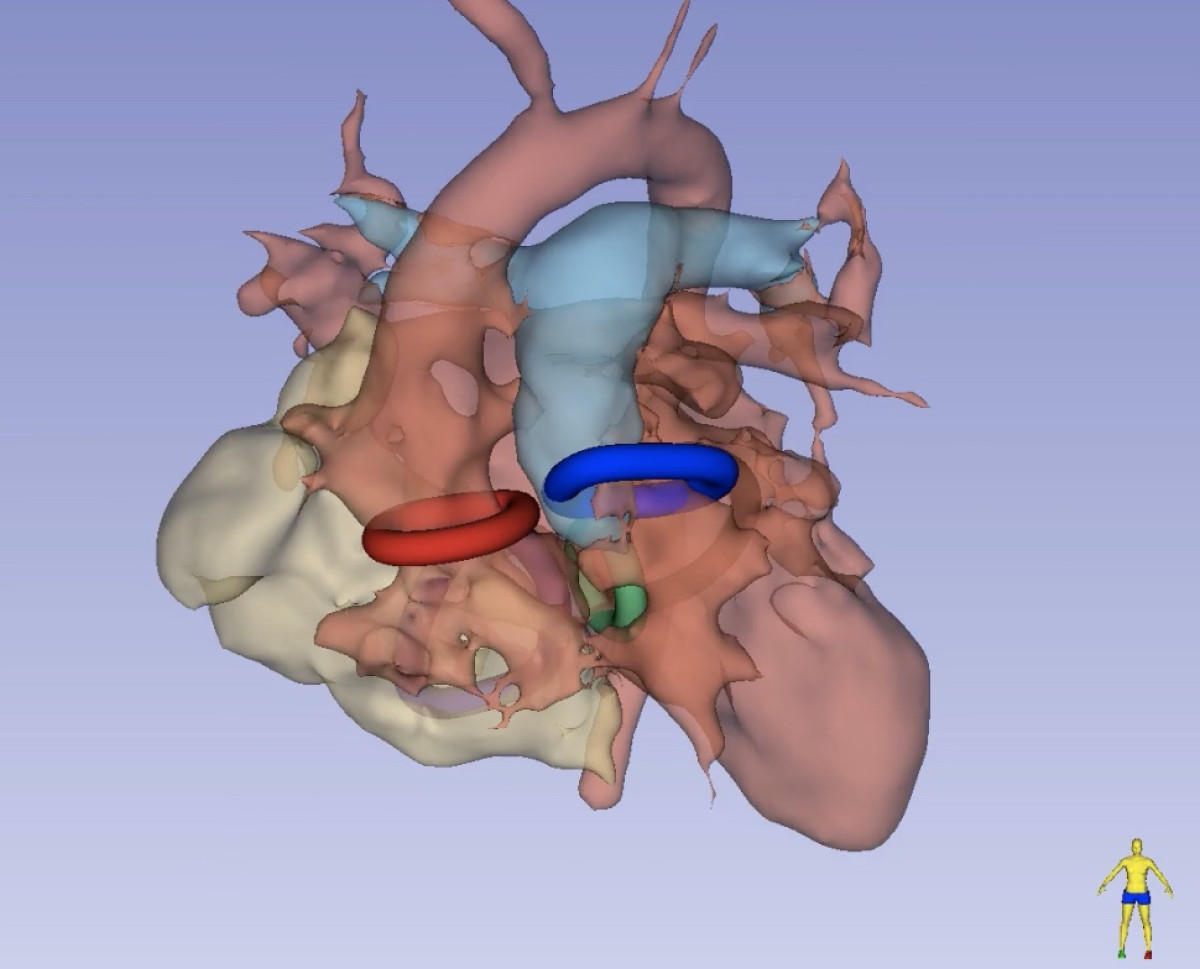
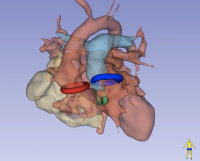
Modeling Intracranial Aneurysm Disease and Management
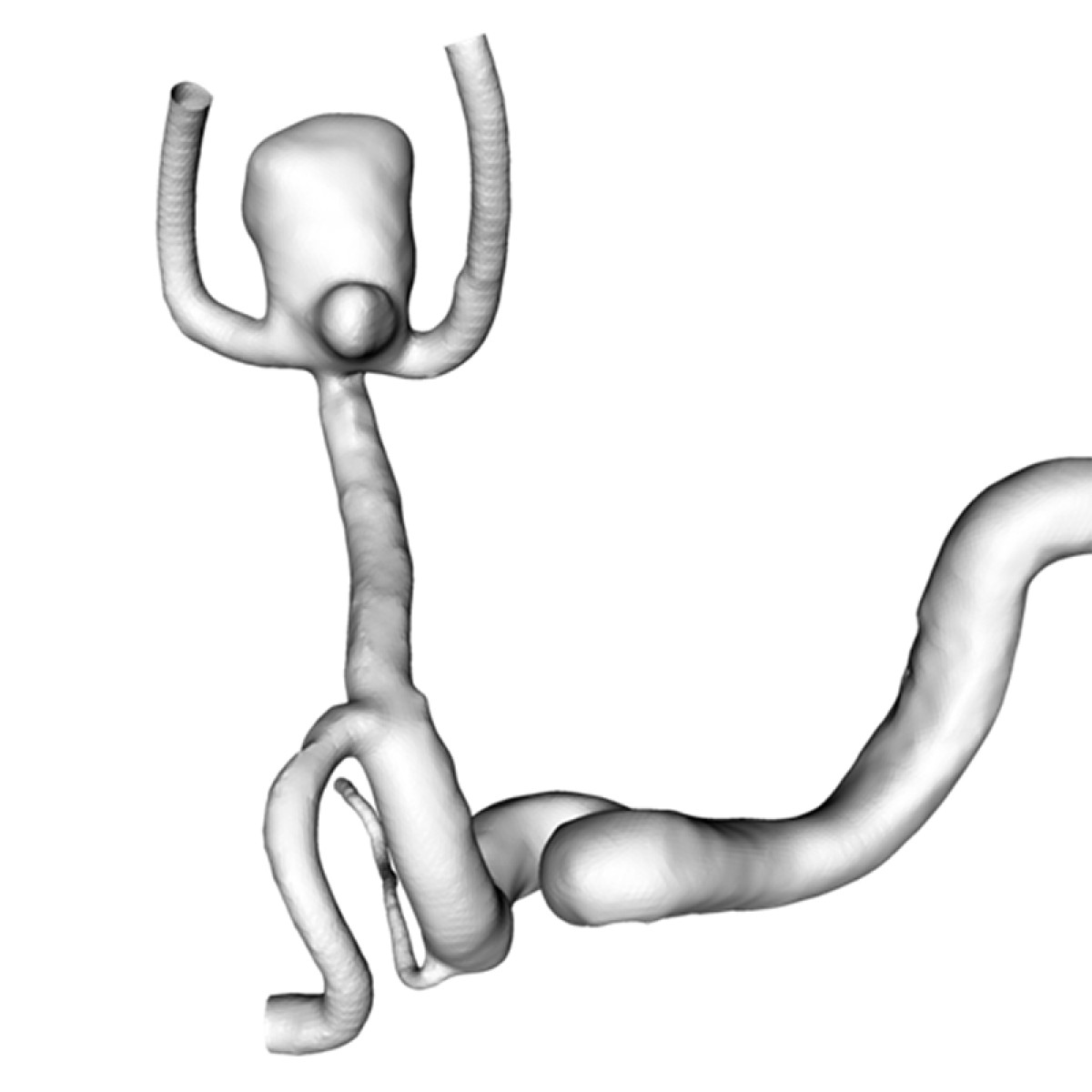
Intracranial Aneurysms are deformations of cerebral vessels that form, progress and may rupture exposing patients to severe disabilities or death. Multiple factors interact at each stage of the disease to result in lesions' initiation and progression. Our research group federates, harmonizes and standardizes information collection from different sources globally. Genetic, biological, imaging and clinical data are analyzed to progressively refine a biomechanical disease model and a statistical phenomenological mode. Both models are used to simulate cerebro-vascular remodeling and offer decision support. In particular, the lab focuses on establishing shape characteristics as a biomarker of disease stage and identifying biomarkers of aneurysm wall stability.
Apply now

Health Informatics for Innovation, Integration, Implementation and Impact
At the core of the development of digital health, biomedical informatics plays a key role in the implementation, integration and evaluation of innovative digital methods and tools. The HI5lab (Health Informatics for Innovation, Integration, Implementation and Impact) aims at connecting these multiple dimensions with the ultimate ambition to demonstrate the impact of eHealth on the health of individuals and populations. The HI5lab is located at Campus Biotech, connected to the various expertise domains such as global health, medical information science, citizen cyberscience, bioinformatics, affective and cognitive sciences. It is also connected to global actors such as WHO, ITU, various UN agencies and NGOs from the International Geneva.
Apply now
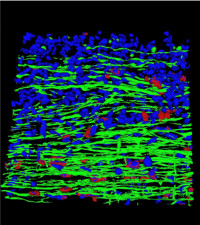
Computational pathology
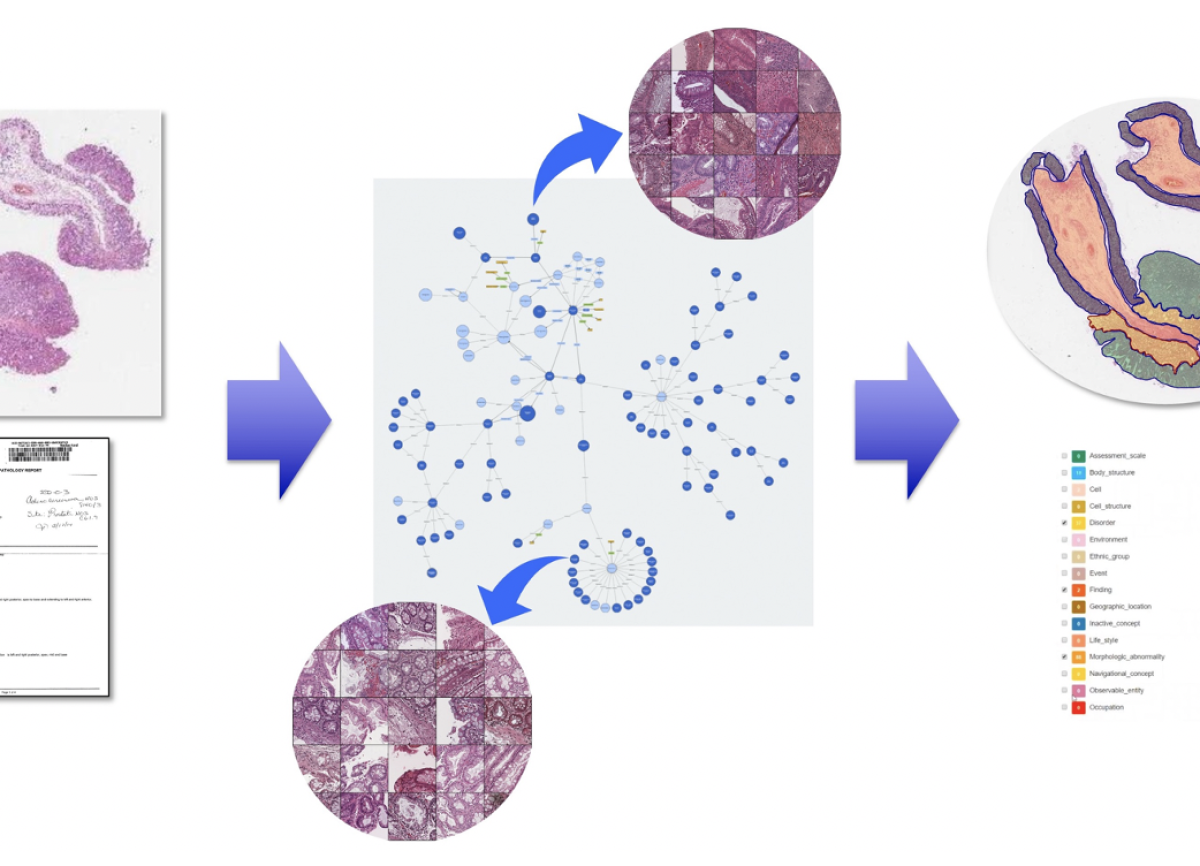
The Computational Pathology Laboratory within the division of clinical pathology is dedicated to advancing the diagnostic and research capabilities of pathology through cutting-edge image analysis algorithms and data integration techniques. Our laboratory combines expertise in pathology, computer science, and data analytics to provide innovative solutions for both clinical applications and research projects by developing and implementing machine learning and AI algorithms to analyze digital pathology images, supporting diagnostic tasks, and conducting research to discover novel biomarkers and predictive models by integrating pathology data with other data modalities.
Apply now
Pharmaco-omics and precision medicine
Our research group's work focuses on investigating genetic determinants of variability in therapeutic response (pharmacogenomics). We study gene-environment-pathology interactions, especially at the pharmacokinetic level, including drug metabolism and transport. Phenoconversion, the discordance between predicted and actual phenotypes, is assessed using the Geneva cocktail. We evaluate genetic vulnerability to drug interactions and develop in vitro, in vivo, and in silico tools to measure and predict drug response. A key focus is optimizing PBPK modeling for personalized prescriptions and progressively integrating phenotypic and pharmacodynamic measures, with future use of machine learning algorithms.
Apply now

Precision oncology
Incorporating clinical biomarkers into oncology treatment strategies allows healthcare professionals to select the most appropriate therapies for individual patients, taking into account their unique genetic and molecular profiles, and thus significantly improving treatment outcomes and reducing the likelihood of unnecessary or ineffective interventions. In our group, we leverage various data sources including pathology images, radiology, clinical and molecular data, to develop actionable biomarkers for oncological treatment. Our goal is to utilize large scale multimodal data and provide machine learning driven approaches that can better guide clinical decisions and benefit oncology patients.
Apply now
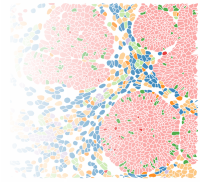
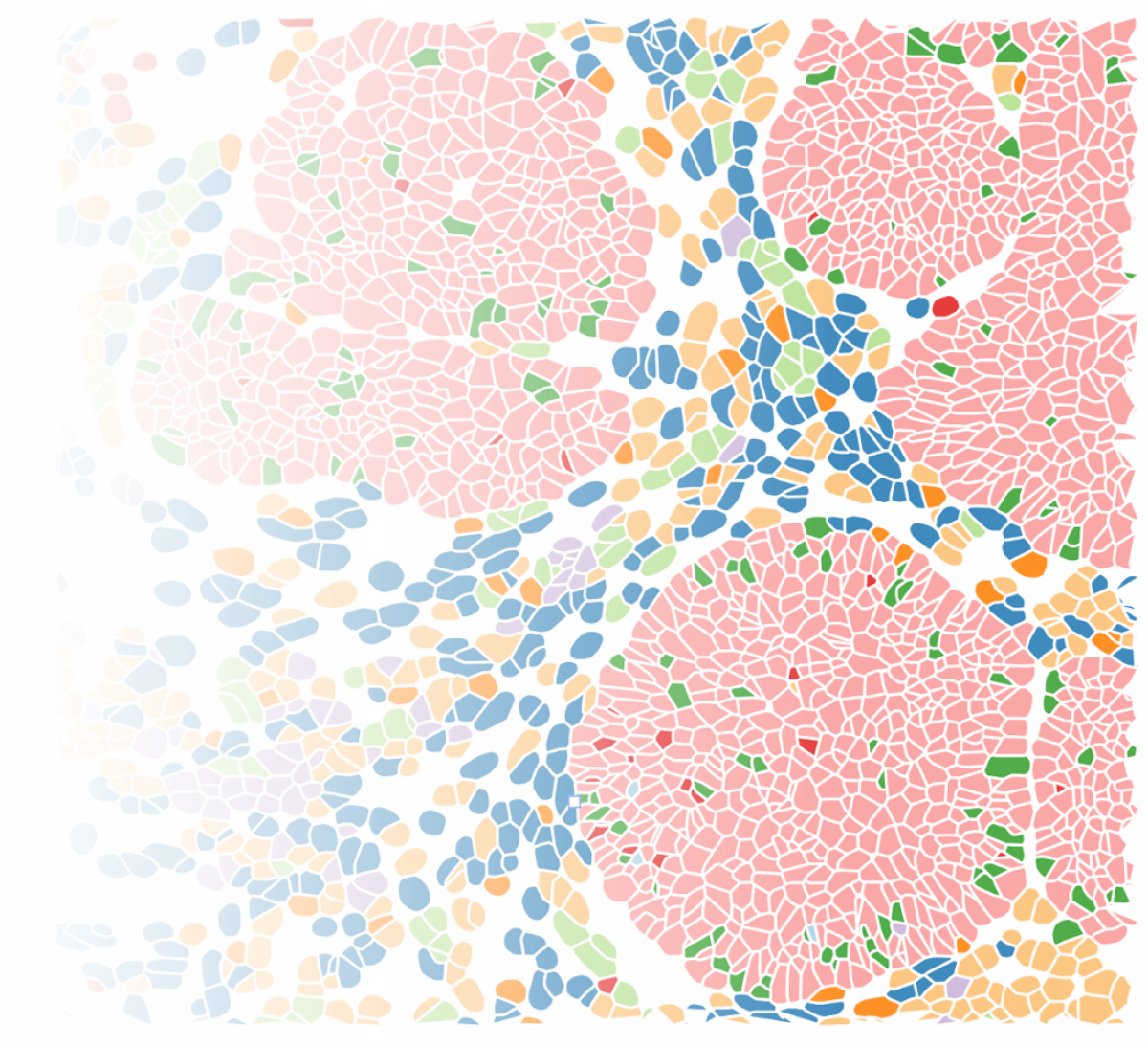
Cancer Systems Immunology
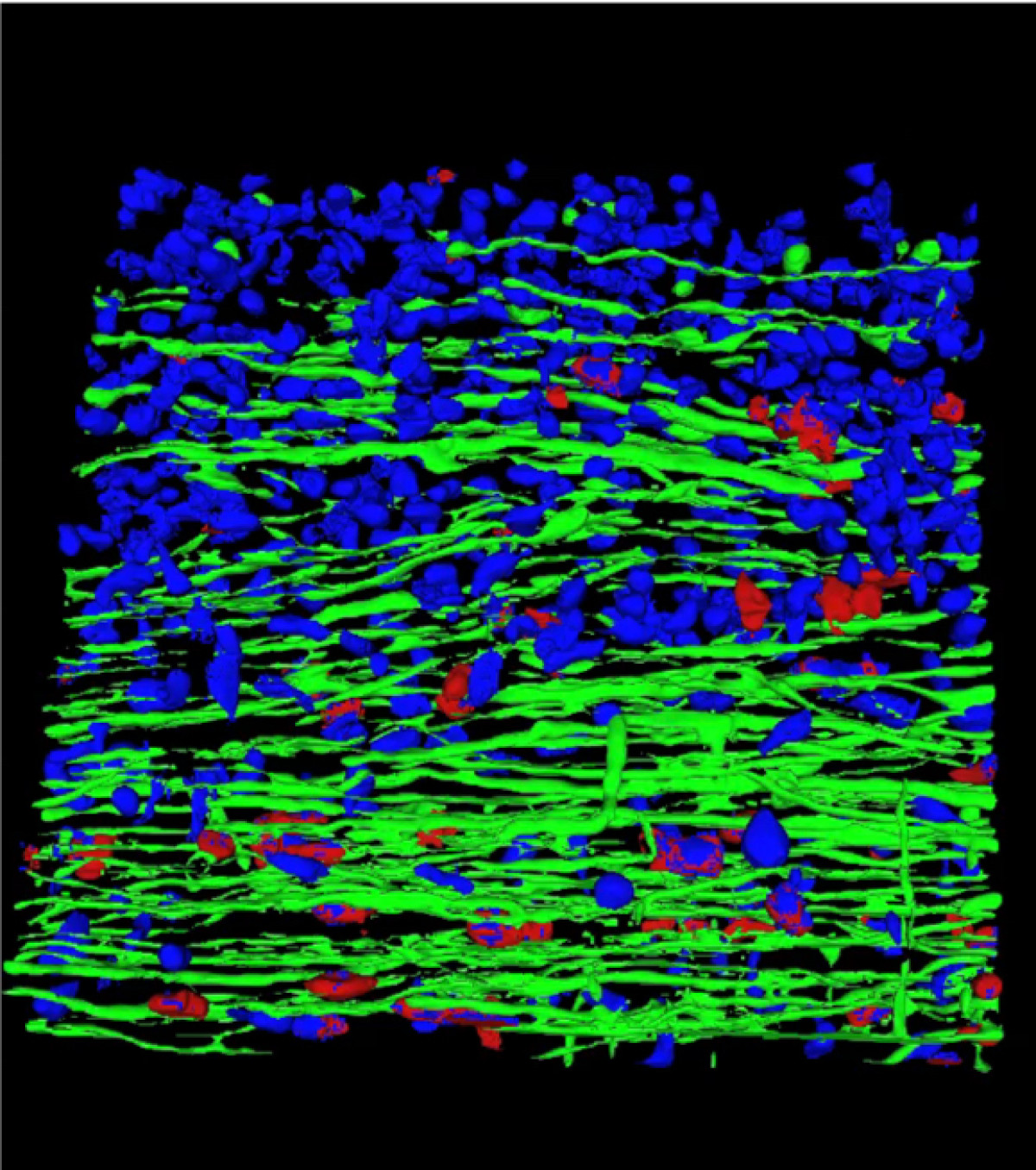
Single-cell and spatial omics technologies are opening new avenues for understanding disease mechanisms at cellular resolution, offering new insights into why diseases and their responses to therapy vary across patients. Our lab develops computational and statistical methods to analyze high-throughput single-cell and spatial omics data. We aim to understand how cell states and cell–cell interactions shape disease trajectories and influence treatment outcomes, with a focus on cancer and immune system dysregulation.
Apply now
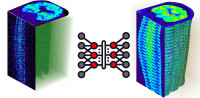
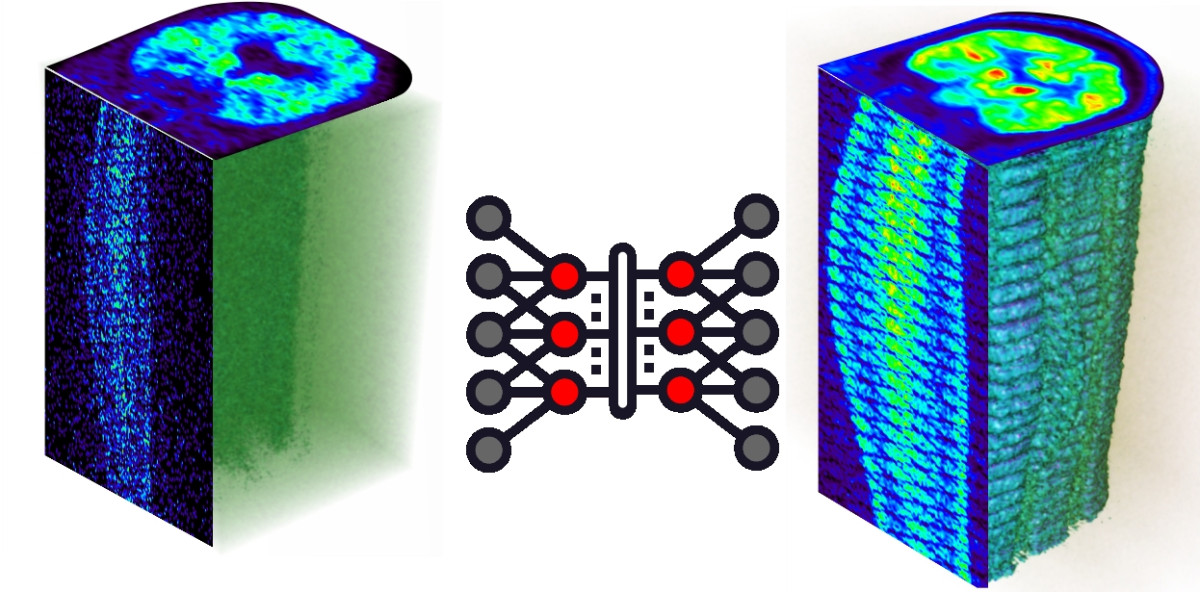
Artificial intelligence in multimodality medical imaging
The focus of our research lies in the application of artificial intelligence in multimodality medical imaging. Our group has assumed a leading role in Switzerland and became internationally recognized for excellence in medical imaging research with multimodality imaging being a focus for its activities. The group gained international recognition for contributions to cutting-edge interdisciplinary biomedical research and clinical diagnosis including the development and validation of new image correction and reconstruction techniques to realize the full potential of quantitative imaging in preclinical and clinical hybrid imaging (PET/CT and PET/MRI) as well as the development of computational modelling and radiation dosimetry algorithms.
Apply now

GIRAPH (geographic information for research and analysis in public health)
We are a diverse group of scientists of different experience and backgrounds, with a shared focus on understanding how spatial, environmental and health data can be combined to provide insights into disease mechanisms and etiologies.
Apply now
Augmented Magnetic Resonance Imaging
Magnetic resonance imaging (MRI) is currently one of the most powerful non-invasive tools to detect and diagnose diseases as well to monitor effect of treatment. Due to the absence of a significant hazard for the human body and to the progress of the technology, the number of images per MRI exam has grown exponentially in the last year. MRI has moved from a purely morphological assessment toward a functional evaluation. This evolution presents a significant challenge related to the large information to process. Located in the Radiology division at the interface of the physics, IT and medicine, our group is specialized in the improvement of the MRI workflow from the image acquisition to the image analysis and clinical validation with a special focus on cardiac and renal diseases.
Apply now

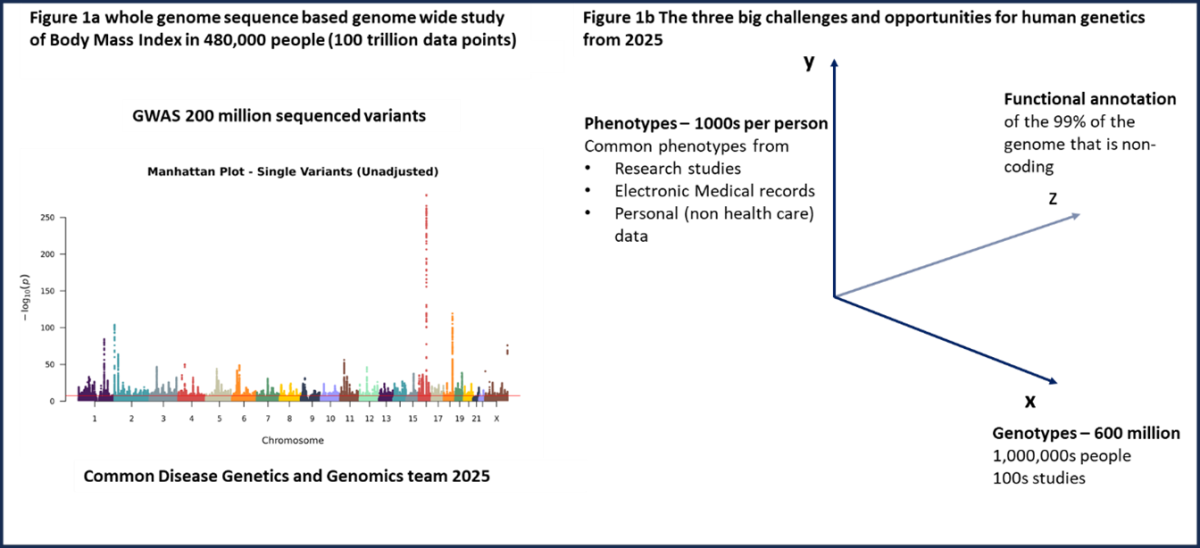
Common disease genomics in an era of whole genome sequencing at scale
The team study the genetics and genomics of common metabolic disease, focusing on both types of diabetes, obesity and related conditions including obesity associated cancers. To do so, we use a combination of genetic analysis, large international and local datasets and clinical studies involving patients and clinicians. To achieve our goals, we adopt an “industry standard” way of working as a team, using an “Agile” approach to project planning. Local datasets we work with will include a new Hospital biobank, “BioHUG” that will incorporate DNA samples and electronic medical records from >20,000 patients in Geneva. There has never been a more exciting time to study common disease using human genetics. This excitement stems from the advances in technology that means we now work with whole genome sequence (WGS) data in 100’000s of people (based on short read sequencing) or 10,000s of people (based on long read sequencing). This use of sequence level data in the context of common disease is still in its infancy but, for the first time, provides us with the ability to identify rarer variants with relatively large effects on common phenotypes across the whole human genome. The discovery of such alleles will facilitate greater understanding of biology, disease and drug development, through more direct functional and clinical studies, and more opportunities for personalised medicine. The advances in genome sequencing are occurring in parallel to two other broad major advances. First, those in functional genomics, where technologies such as single cell RNA sequencing and chromatin accessibility assays means we will have a much greater understanding of the role of the non-coding genome. Second, in the linkage of human genetic and genomic information to routinely collected medical and health information. These 3 big challenges – genotypes, phenotypes and functional information are illustrated as a notional 3 axis graph in Fig 1b.
Apply now

Social and lifecourse epidemiology
Pre Silvia Stringhini is Head of the Population Epidemiology Unit of the Department of Primary Care Medicine at the Geneva University Hospital as well as Assistant Professor at Department of Community Health and Medicine of the University of Geneva. She holds a Master's Degree in Global Health and a PhD in Public Health and Epidemiology. Her main areas of research are social inequalities in chronic diseases and aging, the role of health behaviors in the genesis of social inequalities in health, the biological consequences of social inequalities, the role of environmental factors in social inequalities in health. During the COVID-19 crises, the activities of the Unit have been mostly devoted to the population surveillance of SARS-COV-2, and of the health consequences of the pandemic.
Apply now